Visualizing Biomolecules II: Electron Microscopy

In my last post I talked a lot about how we use x-ray diffraction patterns to calculate the shape of especially tiny things (like biologically relevant molecules). But I segued into it with something that was not an image generated by analyzing x-ray diffraction patterns: I stared by using this image, which is an electron micrograph of a DNA fiber.

Electron microscopy is, at base, more similar to light microscopy than it is to x-ray diffraction. For more on how electron microscopy works – and for a brief synopsis on when diffraction is more useful and when electron microscopy is more useful, keep reading.
Last time I went to great lengths to talk about how the limits of the human eye, and the inverse relationship between wavelength and energy in the visible spectrum, means we can’t just photograph exceptionally tiny things like molecules. One way to get around that is to use high-energy, short-wavelength light: x-rays. But x-rays tend to break apart molecules, so if you use x-rays to “look at” a molecule, you have to use a crystal. So, somehow, what we’d really like is something with a small wavelength – like an x-ray – and a low energy – like visible light.
It turns out, that thing is an electron. Just like a photon, an electron has both wave-like and particle-like properties. Just like a photon, electron wavelength and electron energy are inversely correlated. But for a given energy, the electron’s wavelength will be much smaller than the photon’s wavelength. That means we can use electrons instead of photons and get much better resolution in the resulting images, using processes that are pretty much directly analogous to traditional light microscopy.

Of course, the fact that we’re using electrons and not photons means that even if our methods are analogous, the resulting image still isn’t a photograph: there are generally no photons being graphed. Which is why we call them electron micrographs, when we are being nitpicky. But here’s a quick overview of main kinds of electron micrographs:
“Scanning” electron microscopes are special because they ‘scan’ the image by shooting electrons at it bit-by-bit instead of all at the same time. This is roughly equivalent to confocal microscopy. In each case, a very focused beam of electrons (or light) is used to probe a tiny portion of the sample. That beam is then moved piece by piece until it has covered the entire sample, and the resulting image is stitched together by a computer.
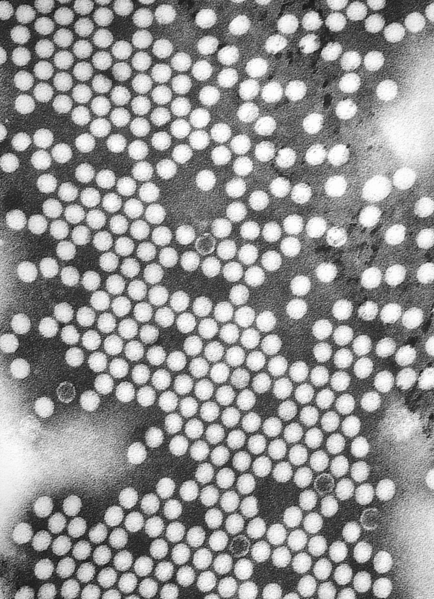
“Transmission” electron microscopes shoot electrons through a sample and see what comes through, rather than seeing what bounces back. Even in light microscopy this is actually fairly common, because very thin biological samples are often transparent or translucent. So, for example, you can shine a light straight through a developing zebrafish embryo or a slide of cells, and see the insides. In a light microscope, different tissue compositions or organelles will diffract different wavelengths of light in different patterns, and you’ll see organs or organelles that way. The same holds true for transmission electron microscopy: different wavelengths of electrons interact with the sample in different ways, and we can chart out parts of the sample by seeing what comes through, where.
The alternative to transmission electron microscopes are, of course, reflection electron microscopes, which measure the reflected scatter patterns of electrons just like you would shine a light on something opaque in order to see it.
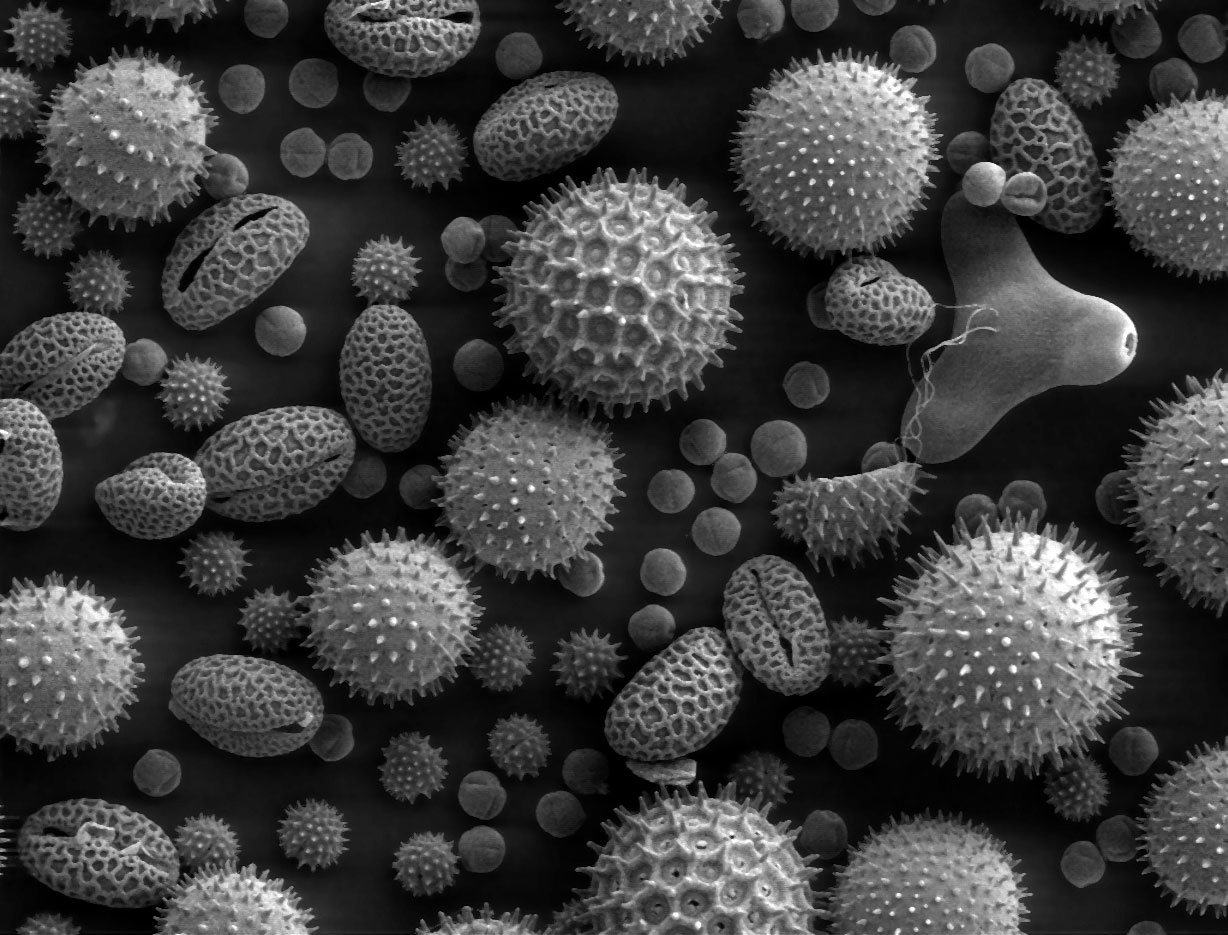
Scanning electron microscopes are great because since you’re only looking at one piece of the image at a time, you can visualize much bigger samples (as much as a few centimeters, which on these scales is ENORMOUS), and the depth of field is fantastic. (As a side note: this is one of the big advantages of confocal microscopy as well.) When you see the electron micrographs of hair or bugs, these are usually scanning electron micrographs.
Transmission electron micrographs have the advantage of a greater resolution, because just like with photons, higher energy (and hence shorter wavelength) electrons pass through samples rather than bouncing back out. Since we’re looking for what passes through, we use shorter wavelengths, and we can get finer-scale images. Images of viruses, protein complexes, and the latest image of DNA are all transmission electron micrographs.
Which, perhaps, brings up the question of: why do diffraction experiments at all? The amount of work it takes to make a crystal – and not only to make a crystal but to ensure that the shape of the units in the crystal is similar to the shape of a molecule in a cell – is enormous. The amount of work that has to be done to analyze the diffraction patterns is significant as well. Is electron microscopy a panacea – a way to look at the tiny building blocks of life in their native environment, without needing to apply a Fourier transform to everything?
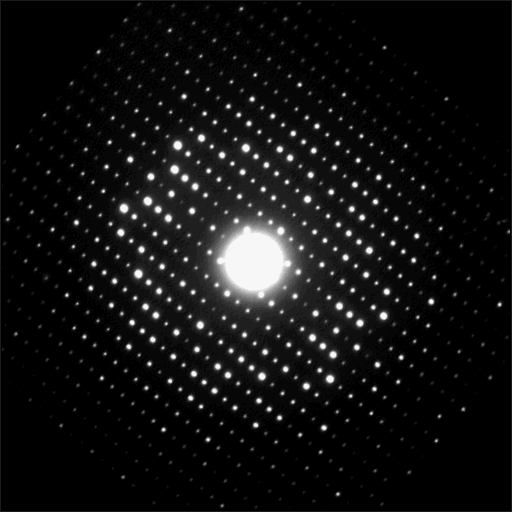

Well, kind of. The fact is, diffraction patterns give you finer-scale resolution than electron micrographs. The shape of DNA was calculated based on x-ray diffraction experiments with much greater resolution than the blurry fiber in the recent breakthrough electron micrograph; and in that blurry fiber you can’t really tell if it’s six or seven strands, or how they’re wrapped around each other. There are people who use electrons instead of x-rays to create diffraction patterns, for basically the same reasons: using a high energy electron and examining diffraction patterns gives you atom-scale information about the shape of a molecule, whereas checking to see how electrons bounce back or pass through a sample cannot.
The fact of the matter, when it comes down to it, is this: electron microscopy is great for asking the question “What does this specific sample look like?”. Crystallography and diffraction patterns are better for asking “What shape does this molecule usually have?”. Being able to examine DNA strands directly in an electron microscope will, hopefully, give us a greater understanding of how sequence and chemical modifications change the shape of a DNA coil. But if, all these years after its initial discovery, we had to rediscover the structure of DNA, we would still be using diffraction experiments to do so.
